Author: Semih Başaran, MD – Clinical Microbiology Resident
Introduction to Pseudomonas aeruginosa Antibiotic Resistance
Pseudomonas aeruginosa antibiotic resistance represents one of the most serious challenges in modern clinical microbiology. This opportunistic Gram-negative pathogen is a leading cause of hospital-acquired infections, particularly in intensive care units. Moreover, its ability to resist multiple antibiotic classes significantly limits therapeutic options.
Over the past decade, the global rise of multidrug-resistant and carbapenem-resistant strains has increased morbidity, mortality and healthcare costs. Therefore, understanding the mechanisms behind Pseudomonas aeruginosa antibiotic resistance is essential for clinicians, microbiologists and antimicrobial stewardship teams.
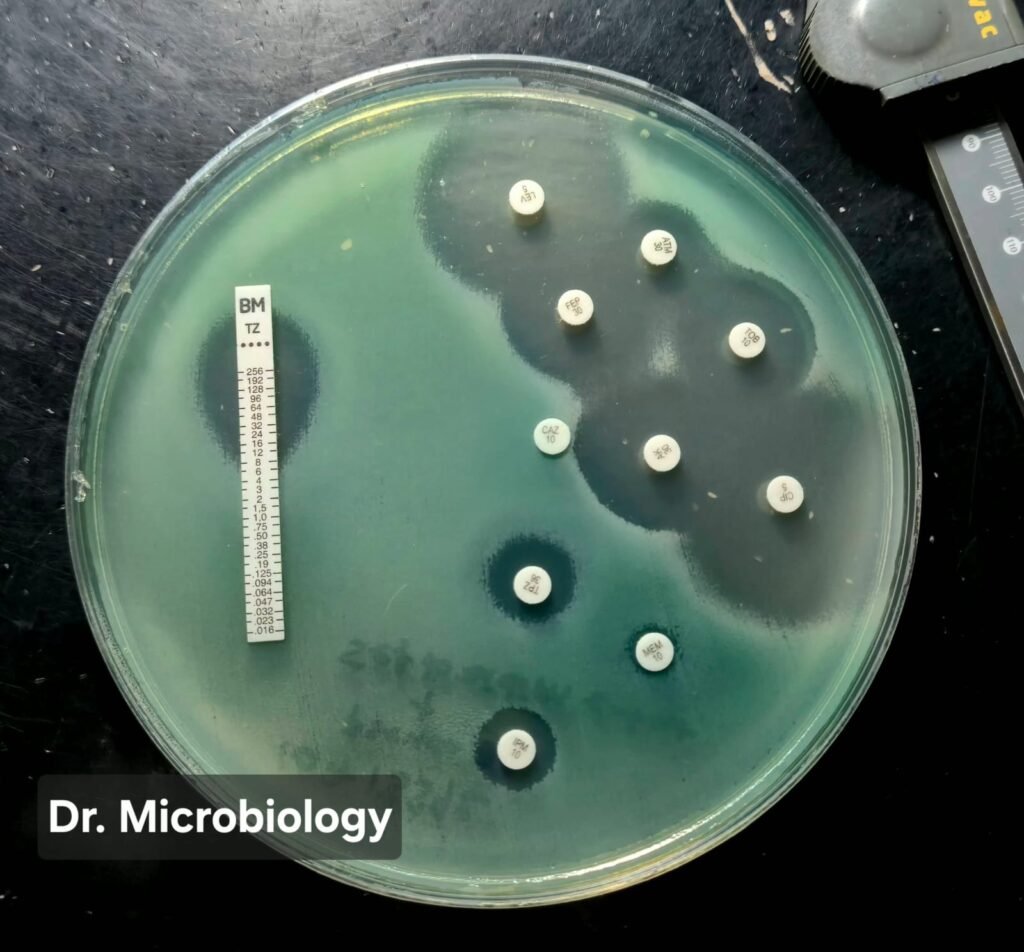
Figure 1. Disk diffusion assay of Pseudomonas aeruginosa performed on Mueller-Hinton agar.
Mechanisms of Pseudomonas aeruginosa Antibiotic Resistance
The remarkable adaptability of P. aeruginosa is due to a combination of intrinsic and acquired resistance mechanisms. Importantly, these mechanisms often act together, producing high-level resistance.
Intrinsic Resistance Mechanisms
Low Outer Membrane Permeability
One of the primary contributors to Pseudomonas aeruginosa antibiotic resistance is its naturally low outer membrane permeability. Compared to Enterobacterales, fewer porin channels allow antibiotic entry. As a result, many β-lactams exhibit reduced activity.
Efflux Pump Systems
In addition, efflux pumps such as MexAB-OprM actively expel antibiotics from the bacterial cell. These systems contribute to resistance against:
- Fluoroquinolones
- β-lactams
- Tetracyclines
- Chloramphenicol
When overexpressed, efflux pumps significantly increase multidrug resistance.
Chromosomal AmpC β-Lactamase
Furthermore, P. aeruginosa possesses an inducible chromosomal AmpC β-lactamase. Hyperproduction of AmpC leads to resistance against penicillins and cephalosporins. Consequently, β-lactam therapy may fail even without acquired resistance genes.
Carbapenem Resistance in Pseudomonas aeruginosa
Carbapenem resistance is particularly concerning. Indeed, carbapenem-resistant Pseudomonas aeruginosa is classified as a critical priority pathogen by the World Health Organization.
Carbapenem resistance develops through several mechanisms:
OprD Porin Loss
Loss or mutation of the OprD porin reduces imipenem uptake. Therefore, isolates may show selective imipenem resistance.
Efflux Pump Overexpression
Increased efflux activity further decreases intracellular carbapenem concentrations.
Carbapenemase Production
Most importantly, acquired carbapenemases such as:
- VIM
- IMP
- NDM
can hydrolyze broad-spectrum β-lactams. Metallo-β-lactamases (MBLs) are especially problematic because they inactivate almost all β-lactams except aztreonam.
Biofilm Formation and Antibiotic Resistance
Another critical factor in Pseudomonas aeruginosa antibiotic resistance is biofilm formation. Biofilms provide a protective environment that reduces antibiotic penetration and promotes persistence.
Within biofilms:
- Antibiotic diffusion is limited
- Bacteria enter a metabolically inactive state
- Host immune responses are reduced
Consequently, biofilm-associated infections such as ventilator-associated pneumonia and catheter infections are difficult to eradicate.
Laboratory Detection of Pseudomonas aeruginosa Antibiotic Resistance
Accurate laboratory detection is essential for guiding therapy.
Disk Diffusion Testing
Disk diffusion remains widely used in routine microbiology laboratories. Zone diameters must be interpreted according to standardized guidelines such as EUCAST or CLSI.
Minimum Inhibitory Concentration (MIC)
Broth microdilution is considered the gold standard for MIC determination. It provides precise susceptibility data.
Phenotypic Carbapenemase Detection
Tests such as:
- Carba NP
- EDTA synergy tests
help detect metallo-β-lactamase production.
Molecular Methods
PCR assays targeting resistance genes (e.g., blaVIM, blaIMP, blaNDM) allow rapid confirmation of carbapenemase production.
Clinical Impact of Pseudomonas aeruginosa Antibiotic Resistance
The clinical consequences of Pseudomonas aeruginosa antibiotic resistance are substantial.
Resistant infections are associated with:
- Increased hospital length of stay
- Higher treatment costs
- Increased mortality
Although newer agents such as ceftolozane-tazobactam offer improved activity, therapy must always be guided by antimicrobial susceptibility testing and local resistance patterns.
Global Epidemiology of Pseudomonas aeruginosa Antibiotic Resistance
The global burden of Pseudomonas aeruginosa antibiotic resistance continues to increase, particularly in tertiary care hospitals and intensive care units. Surveillance data from multiple regions indicate rising rates of carbapenem-resistant Pseudomonas aeruginosa, especially in Southern Europe, the Middle East and parts of Asia.
Moreover, resistance prevalence varies significantly between geographic regions due to differences in antimicrobial stewardship policies, infection control practices and antibiotic consumption patterns. In contrast, countries with strict stewardship programs report comparatively lower resistance rates.
Importantly, carbapenem-resistant Pseudomonas aeruginosa has been associated with outbreaks in ICU settings, burn units and oncology wards. Consequently, continuous surveillance and molecular epidemiology studies are essential to monitor resistance trends and detect emerging high-risk clones.
Therefore, understanding global epidemiological patterns is critical for developing region-specific treatment guidelines and public health strategies aimed at limiting the spread of multidrug-resistant strains.
Why Understanding Pseudomonas aeruginosa Antibiotic Resistance Matters
In summary, Pseudomonas aeruginosa antibiotic resistance results from a complex interplay of intrinsic defenses, acquired enzymes and adaptive mechanisms such as biofilm formation.
Therefore, clinicians must integrate laboratory data, resistance mechanisms and clinical context when selecting therapy.
A strong understanding of these mechanisms ultimately supports better antimicrobial stewardship and improved patient outcomes.
References
1. Lister PD, Wolter DJ, Hanson ND. Antibacterial-resistant Pseudomonas aeruginosa: clinical impact…
👉 https://doi.org/10.1128/CMR.00040-09
2. Poole K. Pseudomonas aeruginosa: resistance to the max…
👉 https://doi.org/10.3389/fmicb.2011.00065
3. World Health Organization. Global priority pathogens list…
👉 https://www.who.int/publications/i/item/WHO-EMP-IAU-2017.12
4. Hancock REW. The Pseudomonas aeruginosa outer membrane…
👉 https://doi.org/10.1128/mr.48.1.1-37.1984
5. Li XZ, Plésiat P, Nikaido H. The challenge of efflux-mediated antibiotic resistance…
👉 https://doi.org/10.1128/CMR.00117-14
6. Cabot G, Zamorano L, Moyà B, et al. Evolution of Pseudomonas aeruginosa antimicrobial resistance…
👉 https://doi.org/10.1093/jac/dkr143
7.Quale J, Bratu S, Gupta J, Landman D. Interplay of efflux, ampC, oprD…
👉 https://doi.org/10.1128/AAC.50.5.1633-1641.2006
8. Morita Y, Tomida J, Kawamura Y. MexXY multidrug efflux system…
👉 https://doi.org/10.3389/fmicb.2012.00408
9. Queenan AM, Bush K. Carbapenemases: the versatile β-lactamases…
👉 https://doi.org/10.1128/CMR.00001-07
10. Hall CW, Mah TF. Molecular mechanisms of biofilm-based antibiotic resistance…
👉 https://doi.org/10.1093/femsre/fux010
11. Bjarnsholt T. The role of bacterial biofilms in chronic infections…
👉 https://doi.org/10.1111/apm.12099
12. CLSI. Performance Standards for Antimicrobial Susceptibility Testing, M100…
👉 https://clsi.org/standards/products/microbiology/documents/m100/
13. EUCAST. Clinical Breakpoints Tables…
👉 https://www.eucast.org/clinical_breakpoints/
14. Nordmann P, Poirel L, Dortet L. Rapid detection of carbapenemase-producing bacteria…
👉 https://doi.org/10.1128/JCM.06026-11
15. Mendes RE, Bell JM, Turnidge JD, et al. Molecular epidemiology of β-lactamase genes…
👉 https://doi.org/10.1128/AAC.48.3.1193-1201.2004
16. Bassetti M, Vena A, Croxatto A, Righi E, Guery B. Treatment of infections caused by MDR Pseudomonas…
👉 https://doi.org/10.1080/14787210.2018.1496722
